Making a shiny app to visualize brain cell type gene expression
Attention conservation notice: A post-mortem of a small side project that is probably not interesting to you unless you're interested in molecular neuroscience.
This weekend I put together an R/Shiny app to visualize brain cell type gene expression patterns from 5 different public data sets. Here it is. Putting together a Shiny application turned out to be way easier than expected -- I had something public within 3 hours, and most of the rest of my time on the project (for a total of ~ 10 hours?) was spent on cleaning the data on the back end to get it into a presentable format for the website.
What is the actual project? The goal is to visualize gene expression in different brain cell types. This is important because many disease-relevant genes are only expressed in one brain cell type but not others, and figuring this out can be critical to learning about the etiology of that disease.
There's already a widely-used web app that does this for two data sets, but since this data is pretty noisy and there are subtle but important differences in the data collection processes, I figured that it'd be helpful to allow people to quickly query other data sets as well.
As an example, the gene that causes Huntington's disease has the symbol HTT. (I say "cause" because variability in the number of repeat regions in this gene correlate almost perfectly with the risk of Huntington's disease development and disease onset.) People usually discuss neurons when it comes to Huntington's disease, and while this might be pathologically valid, by analyzing the data sets I've assembled you can see that this gene is expressed across a large number of brain cell types. This raises the question of why -- and/or if -- variation in its number of repeats only causes pathology in neurons.
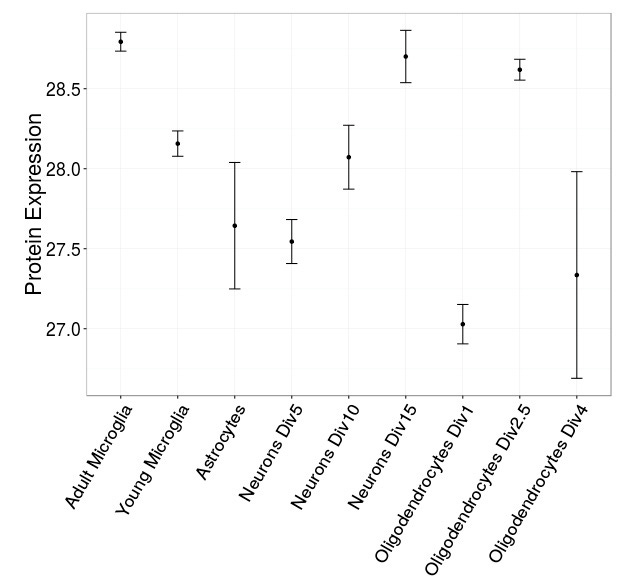
Here's another link to the web app. If you get a chance to check it out, please let me know if you encounter are any problems, and please share if you find it helpful.
References
Aziz NA, Jurgens CK, Landwehrmeyer GB, et al, et al. Normal and mutant HTT interact to affect clinical severity and progression in Huntington disease. Neurology. 2009;73(16):1280-5.
Huang B, Wei W, Wang G, et al. Mutant huntingtin downregulates myelin regulatory factor-mediated myelin gene expression and affects mature oligodendrocytes. Neuron. 2015;85(6):1212-26.

