Question #18: Modeling differential equations in neurochemistry
Modeling is at the crux of what makes science iterative. Andrews and Arkin's 2006 review (here) explains many of the major equations in modeling the reactions of chemical species in cells.
The most common model is the kinetic ordinary differential equation (ODE). It assumes that 1) concentrations of molecules are continuous (even though individual molecules are of course discrete), 2) reactions occur in a homogeneous region, and 3) reactions occur deterministically. The molecules in the reaction are described as a set of differential equations with one equation per chemical. Typically, these equations are solved numerically via computers, although this process is non-trivial.
One of the other common models accounts for the subcellular spatial localization of proteins and reaction components. The concentration at each analyzed region is modeled as a point in a vector, and the reaction is analyzed by taking the partial derivative of these vectors with respect to time. Thus it is called a partial differential equation (PDE). PDEs have the same assumptions as ODEs except that the homogeneous region (assumption #2) is of course not necessary. In order to achieve high spatial resolution, the equations require small subvolumes and short time steps. So PDEs are especially intensive computationally.
One of the main applications of modeling PDEs in neuroscience is in calcium wave models. Calcium imaging relies on indicator molecules that change spectral properties upon binding to calcium ions. So by measuring the fluorescence of the system it is possible to determine the quantity of calcium in a given region.
In particular, this is useful for measuring the rapid propagation of calcium waves through networks of astrocytes connected by gap junctions. Hung et al (here) show an example of an astrocytic calcium wave that can turn growth cones to guide growing axons:
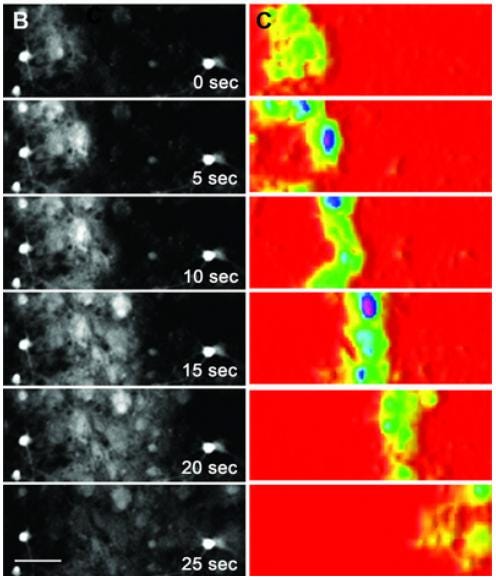
With differential modeling, this type of calcium wave could be fit to a particular model that would help uncover relevant parameters, such as the initial Ca2+ concentration and the Ca2+ flux density amplitude. These parameters can then tell us things about how the astrocytic calcium wave's propagation correlates with other relevant variables, like neuronal activity and axon mobilization.
Inspired by CalTech's Question #18 for cognitive scientists: "Explain how a system of chemical reactions can be represented as ODEs and PDEs. What approximations are involved?"
References
Andrews SA, Arkin AP. 2006 Simulating cell biology. Current Biology. doi:10.1016/j.cub.2006.06.048.
Hung J, Colicos MA (2008) Astrocytic Ca2+ Waves Guide CNS Growth Cones to Remote Regions of Neuronal Activity. PLoS ONE 3(11): e3692. doi:10.1371/journal.pone.0003692


