Modeling the development of the Xenopus spinal cord
There are (at least) two major approaches to building structural models of brain regions: 1) to map all of the connections precisely, or 2) to simulate the development of the connections using validated rules.
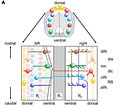
Borisyuk et al.'s study in modeling the developmental of the spinal cord in a Xenopus tadpole is an example of the latter. Here is their general model of the spinal cord that the researchers simulated the development of:

First, the researchers split their model of the spinal cord into subintervals, each containing empty "spaces" that a neuron could "fill." Then, in each of these subintervals, they added the appropriate number of neurons of each type, randomly shuffling their relative positions.
This "appropriate number" of each type of neuron was given by previous experimental evidence, either via antibody staining, anatomical labeling (i.e., injecting horseradish peroxidase), or intracellular labeling with neurobiotin. The empirical distribution of these neuron types along the longitudinal axis is below:

After assigning the positions of the ~2000 neurons, the researchers sampled from an experimental dendrite length distribution to assign dendrites to each of the neurons.
Finally, the researchers used a model for axon generation and growth, which updates the position and orientation of the growth cone at each time step. They optimized the parameters controlling this updating to fit with experimental data for the axons of different cell types.
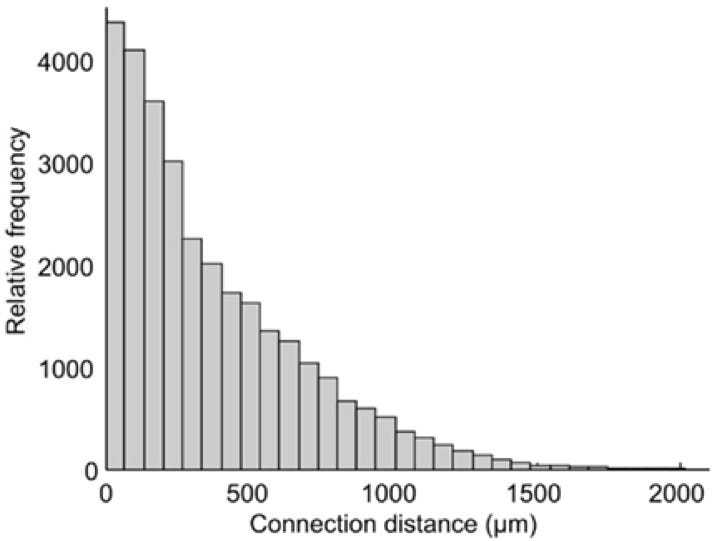
One of the interesting analyses they did with their connectome was to quantify the relative frequency of synapses with various connection distances. As you'd expect and can see below, the relative frequency drops off logarithmically with increasing distance.
Since their developmental model follows stochastic rules, it might be interesting to simulate many connectomes and get a sense of the variability between simulations. This might help us to answer the question, how robust are different sets of developmental rules?
As computational power and the ease of programming continues to expand, such an extension might become relatively easy.
Reference
Borisyuk R, al Azad AK, Conte D, Roberts A and Soffe SR (2011) Modeling the connectome of a simple spinal cord. Front. Neuroinform. 5:20. doi: 10.3389/fninf.2011.00020