Notes on segmenting oligodendrocytes in electron microscopy images using Python3 and OpenCV 3.0
Attention Conservation Notice: Not much new here; mostly just notes to myself for future reference. Reading this is unlikely to be a good use of your time, unless you are trying to install OpenCV 3.0 on Python3, in which case, I tremble with sympathy. (In all seriousness, installing it took me about an hour, but I made some trivial mistakes and it could be way quicker.)
Installing OpenCV for Python3
First of all, know that downloading and installing OpenCV (the CV stands for "Computer Vision") for Python on a MacBook Pro can be pretty time consuming; one commenter I saw online called it "a rite of passage." After failing to successfully download it for Python 2.7 for about half hour, I eventually decide to use this opportunity to upgrade to Python3.
Installing Python3 is easy:
[code]brew install python3[/code]
I then followed Luis González's useful tutorial for installing OpenCV as a module for Python3. I still ran into one problem, which is that I had a different version of Python. My recommendation here is to manually cd to those directories in your terminal to make sure that the files exist as they are meant to, and then copy and paste the paths into your Cmake GUI. Here is my successful configuration:

Specifically, if you are getting the error
[code]fatal error: 'Python.h' file not found #include <Python.h>[/code]
(perhaps at 98% completion!), make sure to check that your $PYTHON3_INCLUDE_DIR is set properly; mine had a typo.
How to identify oligodendrocytes in electron microscopy images
Once I had OpenCV downloaded, I was able to begin actually analyzing some EM images. I'm interested in being able to distinguish oligodendrocytes on electron microscopy from other brain cell types. It turns out this is a pretty difficult problem, and as far as I know, there is no large database of images from which a substantial training set (e.g., > ~ 200 images or so) could be built.
Instead, we can go based on published features of oligodendrocytes, which have been described previously (e.g., here, here, here, here, and here). As far as my novice understanding goes, these are some key features of oligodendrocytes:
Smooth, round-to-oval outline
Centrally placed, round-to-oval nuclei, (with "nucleoli occasionally seen on the plane of section")
Distinct and dark Golgi apparatus in the cytoplasm
Thin processes (when they are visible)
And here are some features distinguishing oligodendrocytes from other cell types:
Astrocytes: a) paler (cytosplasm and nuclei), b) glycogen granules, c) filament bundles (due to GFAP), and d) wider processes (if the processes can be identified)
Neurons: large and pale nuclei
Microglia: more likely to be irregularly shaped, and sometimes have dark, small inclusions
NG2 cells: elongated or bean-shaped nuclei, and can contain long endoplasmic reticulum
Here is a picture of an oligodendrocyte from Alan Peter's helpful website:
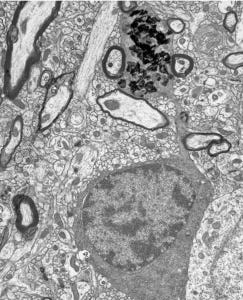
Using OpenCV to parse tissue slices
Here is my code. I tried a few different strategies. My goal is to parse slice out a portion of the image that is recognizable as "the oligodendrocyte," which can be seen by the human eye as the red portion here (also from Alan Peter's website):

1) Canny edge detection. This seems to be "too much" to be useful.

2) Otsu's reduction of a greyscale to a binary image. This is actually not bad; maybe it could be stacked with another method?

3) Blob detection. This is somewhat promising, but unfortunately the blobs detected are pretty small (too small to be the oligodendrocyte, or even the nucleus), and when I try to make them larger, no blobs are detected.

So, this is still a work-in-progress, but I wanted to get some notes up.
Reference
The fine structure of the aging brain. Authors: Alan Peters and Claire Folger Sethares. Boston University School of Medicine, 72 East Newton Street, Boston, MA 02118. Website: www.bu.edu/agingbrain. Supported by the Institute on Aging of the National Institute of Health, grant number P 01-AG 000001.


