The goal of brain preservation is to preserve and protect the information in someone’s brain indefinitely, with the aim of potential revival if future civilizational capacity ever allows it to be feasible and humane.
Since we’re considering a hypothetical outcome in the distant future, a key question is: how can we evaluate the quality of preservation today? We need metrics to avoid wasting time and money on useless rituals.
The first main divide is between people who favor functional vs structural metrics. I tend to be on the side that structural metrics are best. Our hypothetical restoration technology — whether that is via molecular nanotechnology, whole brain emulation, or something else entirely — is likely to be so much better than the technology we have today that trying to test functional read-outs today will not be very dispositive. Instead, I think the best metric we have is to use microscopes and look at the tissue in detail to see whether the structures that seem to be important for memories, personality, and etc are intact.
But that raises the question of which structural metrics one should be evaluating when comparing between possible brain preservation procedures. And this gets us to another divide, between people who strictly favor connectome traceability via contemporary electron microscopy and others, like me, who think that is a great ideal goal to aim for, but is probably not strictly necessary. I'm mostly writing this blog post because I keep encountering this disagreement and I want to make my position clear.
Connectome traceability by contemporary electron microscopy
Let’s first try to understand this position. As far as I can tell, two of the most prominent researchers who have advocated for it are Ken Hayworth and Robert McIntyre.
I always like to try to understand the history of ideas. Let’s jump back to around 2011. Although cryonics was first proposed in the 1960s, and many were bullish on near-term revival at the time, it didn’t happen. Instead, one of the main justifications for cryonics practice was structural — if you cryopreserve brain tissue in the right way and rewarm it, a lot of the structures still look like they are there, albeit damaged.
Mike Darwin’s 2011 blog post, “Does Personal Identity Survive Cryopreservation?”, is pretty clearly the best explanation of this stance. The data relied on perfusion of animals with cryoprotectants, either with high concentration or in vitrification protocols. The structures looked intact but also dehydrated.
There also wasn’t a report of a whole brain that had been cryopreserved, rewarmed, processed, and imaged in such a way that would allow for the connectome — the set of all neural connections — to be traced. In 2011, Ken Hayworth proposed the brain preservation prize, which leveraged cutting edge neural imaging technology with the goal of measuring ultrastructural quality and connectome traceability.
As Ken put it:
We now understand that the true measure of success should be that a procedure preserves the structural connectivity of the neuronal circuits of the brain along with enough molecular level information necessary to infer the functional properties of the neurons and their synaptic connections.1
Around the same time, Sebastian Seung also pointed out the necessity of this test to evaluate cryonics in a TED Talk and his book Connectome.
This test is kind of a binary thing: either a procedure preserves the brain in such a way that the connectome is traceable by contemporary standards or it doesn’t. The initial goal was to evaluate the preservation procedure itself, but similar reasoning also can be applied when considering other types of damage occurring during the brain preservation process, such as agonal, postmortem, or storage-associated damage.
Robert McIntyre and Greg Fahy published a paper in 2015 describing aldehyde-stabilized cryopreservation, a procedure that ultimately won the Brain Preservation Foundation prize for preserving the brain in a way that could be stored long-term with connectome traceability intact.
The pure cryopreservation method used in conventional cryonics did not win the prize, and as far as I know, still has not met this standard. The images I have seen from this method applied to whole brains demonstrate dehydration-associated damage, making them difficult to interpret2:
The comparison of aldehyde-stabilized cryopreservation vs pure cryopreservation is not the purpose of this post. Instead I just want to point out that deciding upon metrics has very real-world consequences, because it can help to compare the quality of preservation between methods, and also help people to decide whether a procedure is worthwhile at all.
This metric has a lot of advantages. It really seems to include all of what contemporary neuroscientific models tell us could be involved in memory storage. Ken has tried to ask other neuroscientists why it wouldn’t be sufficient to preserve engrams, and no compelling counterarguments have been made public, other than general uncertainty about future knowledge and capabilities.
But while it is a great ideal metric, I also think it is too stringent, because it is limited by the capabilities of contemporary imaging modalities. I think we should have more consideration for how reasonable future advances in imaging technology, such as expansion microscopy, might make things easier.
With that in mind, how about this mouthful of a metric instead?:
Intact cell membrane morphology via light microscopy, as a proxy for connectome traceability by hypothetical molecular imaging
This metric relaxes the stringency of flawless electron microscopy-level imaging, but still aims to capture the critical structural information.
The reasoning is that if cell membranes, including dendrites, axons, myelin, synapses, and other important structures, maintain their rough shape and location as seen on light microscopy (i.e. are “intact”), then even if they are degraded on the finer electron microscopy level, future molecular imaging may still enable connectome inference.
I want to explain why, but first I have to give a lot of background. Some ballpark resolution numbers might help to ground us. FIB-SEM, which is the type of electron microscopy used to evaluate the brain preservation prize, has a resolution of around 5-20 nm. Routine light microscopy has a resolution of around 200-500 nm. So FIB-SEM has around 10-100x better resolution than routine light microscopy.
Why focus on cell membrane morphology and not other aspects of morphology, like nuclear shape or organelles? Because it seems to me like cell membrane shape is probably one of the most important parts of the connectome, while organelle shape isn’t as critical for memory information.
Of course, one would also need biomolecular information to annotate the connectome, but biomolecular information is largely redundantly stored in the epigenome, other biomolecules, etc, so it doesn’t seem as essential to evaluate as cell membrane preservation. So that’s why I’m focusing on structures visible under the microscope.
To make it clear that I’m not just accepting “anything”, here are a couple of examples of cases that would or would not meet my criteria of intact cell morphology on light microscopy. The first example is any liquefied brain. If something is liquefied at the macroscale level, it seems obvious that the connectome is lost.
But that’s trivial. For a less trivial example, let’s consider Monroy-Gómez et al 2020, a nice article that looked at how long rabies virus antigens could be detected in mouse brain tissue after death using different techniques.
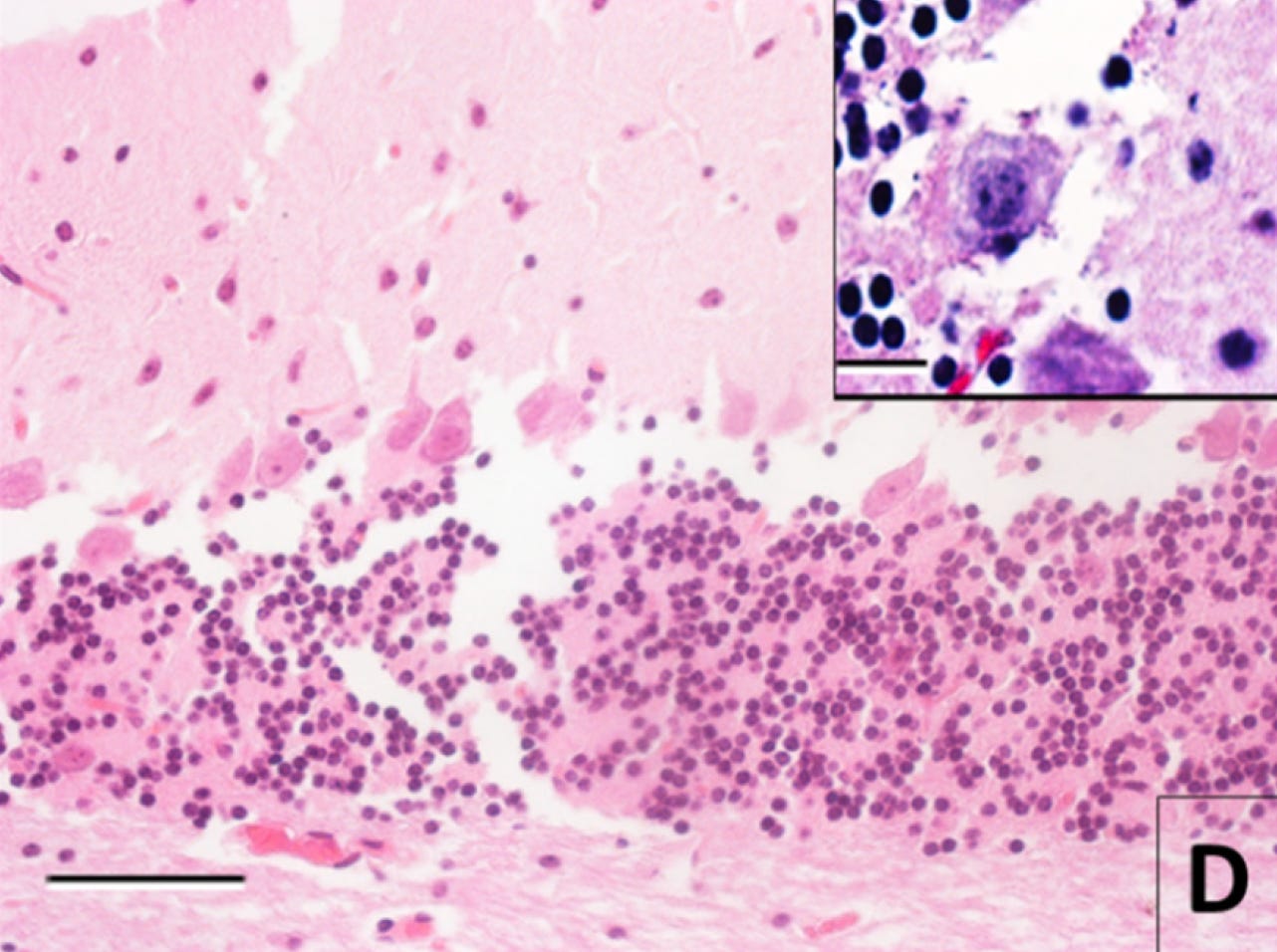
First, here is a control image, an H&E-stained light microscopy image from the cerebellum of a mouse whose brain was perfused immediately after death:
And here is an image after 30 hours of storage at room temperature after death prior to immersion fixation and tissue processing:
To me, the original structure seems not inferable based on this image. With the separations, the cell membrane alterations, the changes to the neuropil, and the fact that some cells from the granule layer are reported to be literally gone. Probably too much damage. Am I sure? Definitely not. But this an example of an image that would not pass my subjective sufficient quality threshold.
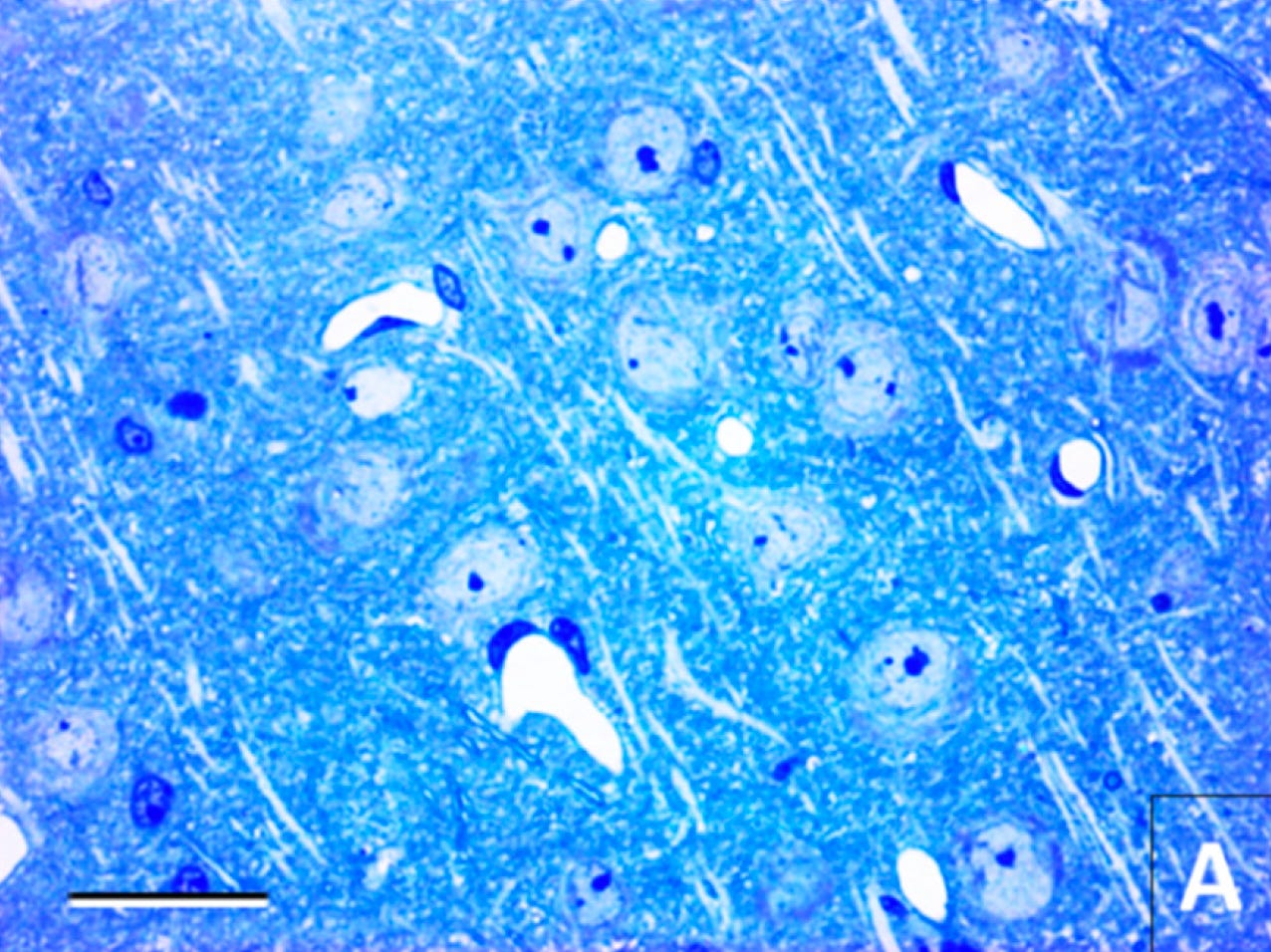
Here is another example from the same paper. This is an image from the cerebral cortex stained with toluidine blue. Here is the control condition of tissue preserved immediately after death. You can see the normal cellular morphology:
And here’s a corresponding image after 30 hours of room temperature storage after death:
In this image, it’s no longer possible to identify the cellular morphology.
What do I mean by molecular imaging?
Molecular imaging refers to imaging techniques that can visualize molecules and biomolecules within cells and tissues, like immunohistochemistry. These techniques allow you to see the location and abundance of specific proteins, nucleic acids, lipids, and other biomolecules.
We may one day have imaging technologies that can map the position of every biomolecule in a tissue sample, or even across the whole brain. This extremely high-resolution molecular imaging could allow inference of cell connectomes and other fine structural details, even if the original tissue morphology is disrupted.
Mechanistically, there’s a lot of structures preventing biomolecular diffusion during the dying process or during the preservation procedure, including cell membranes and the extracellular matrix. And empirically, many types of biomolecules do not seem to immediately diffuse away from their original locations after death. So I think there is surface validity to this idea.
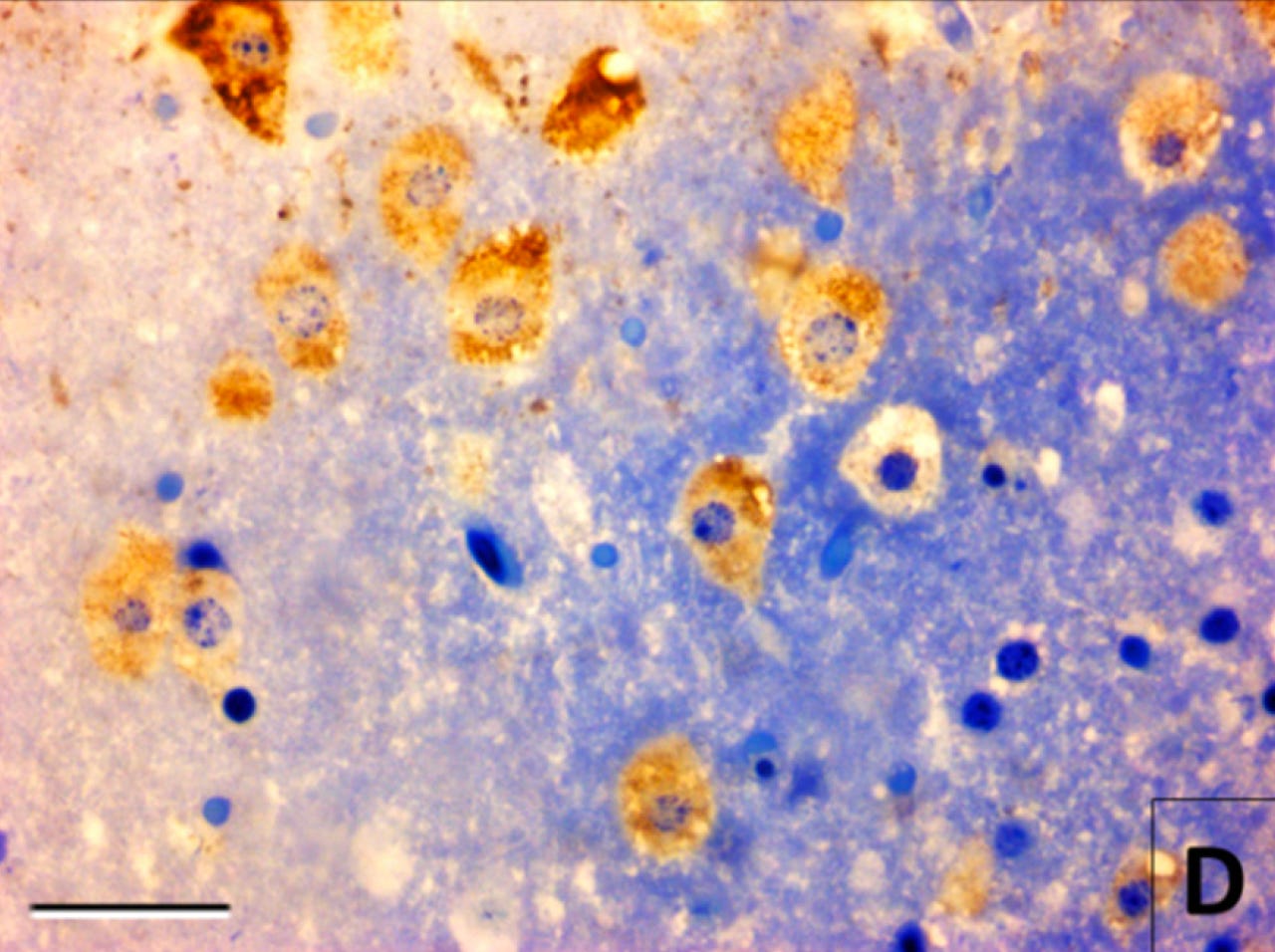
As an example of what I mean by using molecular imaging to infer what morphological imaging cannot, here’s the same tissue in the most recent image, i.e. mouse cerebral cortex preserved at 30 hours after death. However, this time, there is also immunostaining for a rabies antigen, which appears as a brown color:
In this image, suddenly the cell morphology does seem visible. That’s because although the morphological stain — toluidine blue — didn’t bind to biomolecules in the cells specifically enough to allow for contrast, the rabies antigen immunostaining did — at least in the subset of cells infected by the rabies virus.
When I think about future molecular imaging technology, I’m imagining something similar to this, but at the level of single biomolecules, down to their atomic composition if need be. And also allowing for deconvolution of the original state in the case that the biomolecules have diffused from their original places.
What type of biomolecules would we be imaging? Well, every cell has a unique composition of biomolecules. This includes cell surface biomolecules like neurexins and protocadherins that have a variety of gene expression mechanisms to allow for variability between cells, in part so that cellular processes can recognize self vs non-self.
Cells also have a ridiculous number of biomolecules. For example, each neuron has been estimated to have around 50 billion proteins.
Even if there is damage in the dendrites or axons on electron microscopy, so that it’s not obvious how to trace it, it still might be possible to trace that neurite following a molecular imaging process. One would need to image the molecules, identify their most likely cellular origin, and deconvolve the most likely original locations of the cells based on predicting where their constituent molecules diffused from.
What do I mean by deconvolution?
Let me also give an example here. In microscopy images taken postmortem, one often finds that cell membranes are “blurry”. They don’t look sharp like they usually do. For example, in Blair et al 2016, you can see the cell membrane morphology stained for rRNA become more blurry as the postmortem interval extends from 22 hours (left) to 41 hours (right):
Deconvolution is the idea that if you map enough of the biomolecules at a detailed enough level, you can build an inverse diffusion model to infer what the original state most likely was. In this case, to turn the membranes from blurry to sharp again.
Of course this deconvolution procedure will not be perfect, because molecular diffusion is a random process. That’s okay, because the connectome is not static anyway, yet our long-term memories are still stable over a long period of time, suggesting that there is a decent amount of error tolerance in information encoding.
If the brain is liquefied or the local area has undergone too much decomposition, there’s going to be a point where attempting to computationally infer the original state based on mapping the diffusion breakdown products is not going to be possible to a sufficient degree of accuracy. Future molecular imaging technologies will still not be magic.
This is why I think it’s useful to look at light microscopy images. Although it has a lower resolution, light microscopy bounds how much damage could have occurred to neural structures at the electron microscopy level. My guess is that nanometer scale decomposition that might be occurring beyond what is seen via light microscopy, visible only by electron microscopy, could still potentially be inferred by future molecular imaging technology.
While molecular imaging may be able to recover some information from minor degradation, there will still be a threshold of severe degradation beyond which the original state cannot be accurately inferred. Intact cell morphology on light microscopy provides a proxy for whether tissue degradation has surpassed this threshold.
I think biophysically, there has to be some point at which structures are no longer visible by morphological stains, but they would be visible by molecular imaging. What are the structures under the microscope actually made up of? Mostly, they seem to be collections of biomolecular gel-like networks that are strengthened by fixation. When those gels break down, they’re no longer able to be visualized under the microscope. But the biomolecules that make them up don’t immediately disappear. Instead, they dissolve and disperse into the local area.
On the other hand, if cell morphology is absent on light microscopy, I think that is a very bad sign suggesting there is a good chance that the information has been lost (i.e. is no longer inferable via classical physics). Light microscopy also has the advantages of being much cheaper, more accessible, and easier to directly measure biomolecules.
There are plenty of problems with the metric of “intact cell membrane morphology via light microscopy”
Obviously, this metric is far from perfect. It is subjective and depends on the methods used. There are no definitive thresholds or precise cutoffs to determine how much morphological damage is acceptable. Instead, it relies more on an overall gestalt assessment of the integrity of cell shape. This subjectivity makes it difficult to establish the type of objective, binary prize criteria that connectome traceability provides.
Also, the whole molecular imaging idea is an additional area of uncertainty in the brain preservation project. We cannot say for sure whether it will work. The ideal goal — if it is possible — would be to achieve connectome traceability as seen by electron microscopy today, and thereby avoid this whole discussion.
Finally, relying on an imperfect proxy metric risks complacency in pushing for further advances. Accepting intact morphology as “good enough” could reduce incentives to achieve the ideal of connectome traceability.
Despite these problems, this framework is my best attempt to make sense of a complex issue with the information I have available today. Refining this metric to be more quantifiable and definitive is an important goal of mine for the future. The potential for complacency could be mitigated by continually measuring the effectiveness of preservation techniques and not being satisfied with the status quo.
Summary
It seems right to aim for verifiable connectome traceability as seen on electron microscopy as the ideal goal of structural brain preservation. If we have achieved this goal, then I think there is a clear case that the information required for engrams has been preserved, unless there is something very wrong with our theories of neuroscience. The only method that has met this bar is aldehyde-stabilized cryopreservation under ideal laboratory conditions. But, it is a hard bar to reach in most practical cases, where agonal states are severe, perfusion is difficult, and/or the postmortem interval has been too long.
So, for people dying today who can’t access that method — either due to it not being available in their area, cost, brain tissue degradation before the procedure can start, or any other reason — I think it’s also reasonable to use procedures that meet the criteria of intact cell membrane morphology on light microscopy, as a proxy for connectome integrity by a hypothetical molecular scan.
Which brain preservation methods meet this criteria? I have some opinions, but I’m planning to do some more research and make sure my reasoning is sound before I write publicly about them. In the meantime, I just want to make it clear what criteria I currently think should be used: (a) as the “gold standard”, connectome traceability by contemporary electron microscopy, but if that’s not achievable, then (b) intact cell morphology via light microscopy also seems like it would have a reasonable chance of being sufficient.
When I use the term connectome, I’m using it in the sense of also including molecular information.
However, I’m also aware that more recent data using vitrification approaches due to be published is reported to look better.








Makes me think of when there's revival of the dead. They'll freeze upon waking.